| Home | Syllabus | Schedule | Lecture Notes | Extras | Glossary |

| Home | Syllabus | Schedule | Lecture Notes | Extras | Glossary |
We began by reminding everyone that Exam 3 is scheduled for Tuesday, November 12. The developing Study Guide is posted. The study guide will be "frozen" on Friday, November 8.
Our discussion of developmental genetics will be almost entirely confined to Drosophila, as the work there is very complete. Many of the basic principles discovered in Drosophila research apply broadly to development in other animals, including humans.
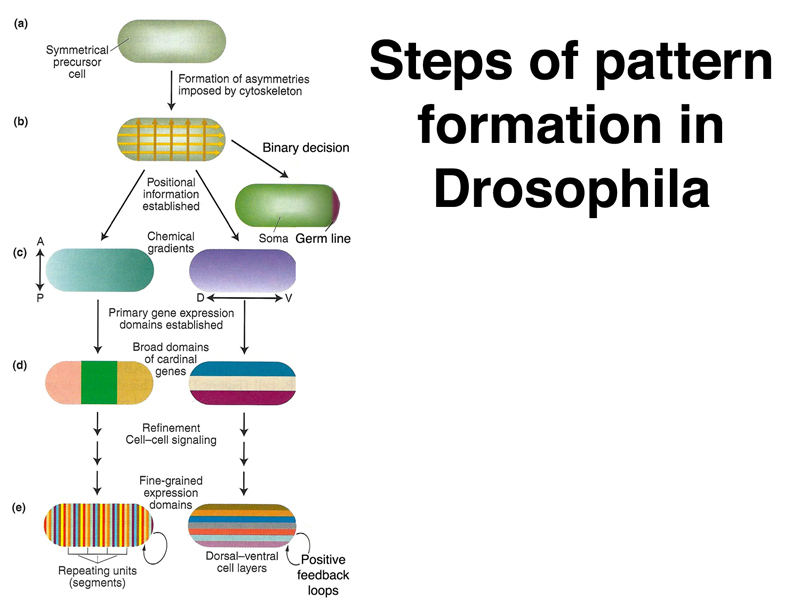
As shown in the overview below, we can think of development in Drosophila as assigning a precise location to each part of the embryo through an anterior-posterior and dorsal-ventral positional code. This positional information, which is used by cells to decide their fate, is successively refined using sets of genes with different roles. Some of these genes are used only in embryonic development and are not expressed once embryonic development is complete.
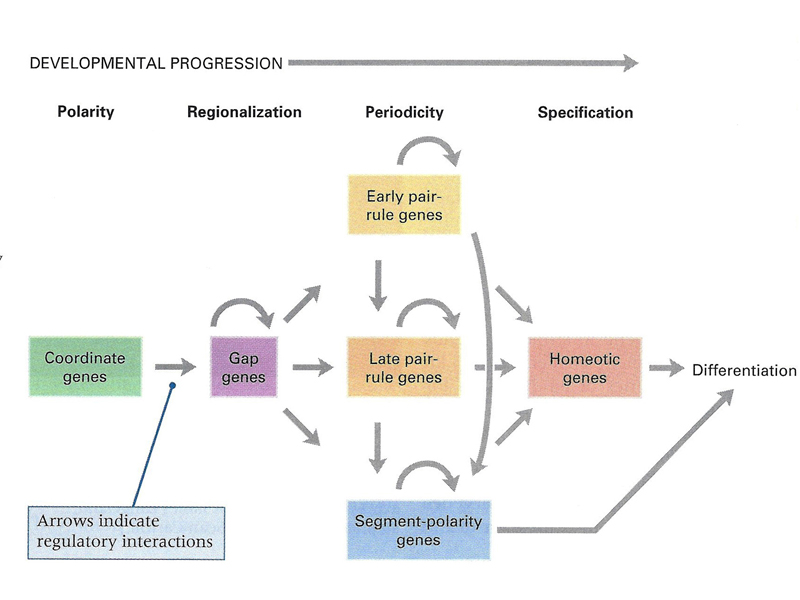
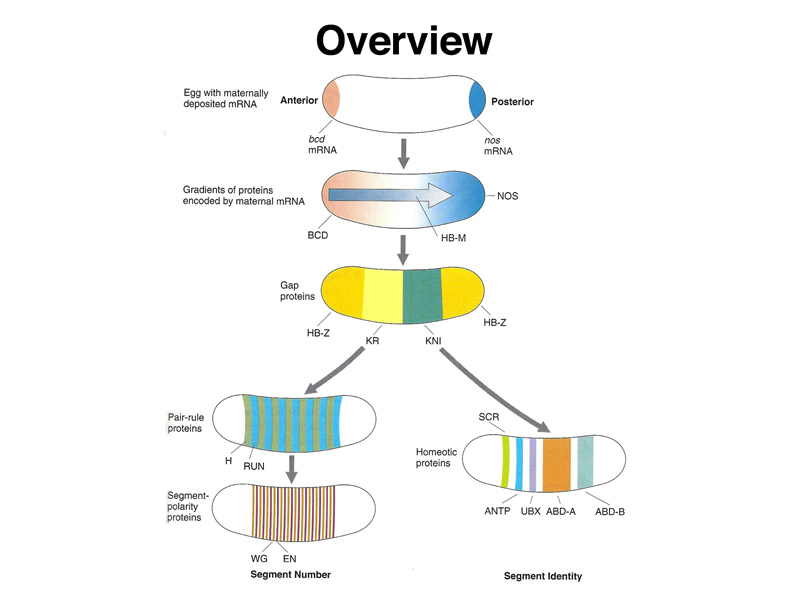
The overview below is more precise, naming some of the genes involved. There is a set of coordinate genes expressed maternally that build positional information into the egg. The coordinate genes establish gradients of morphogens within the egg that are interpreted by the gap genes, the first of the zygotic developmental genes to be expressed. Gap genes are expressed in broad regions spanning a number of adjacent future segments. The pair-rule genes are then expressed in a pattern of alternating future segments, controlling the expression of segment-polarity genes that define portions of individual segments. Meanwhile, homeotic genes are expressed in a pattern that confers a specific identity on each segment.
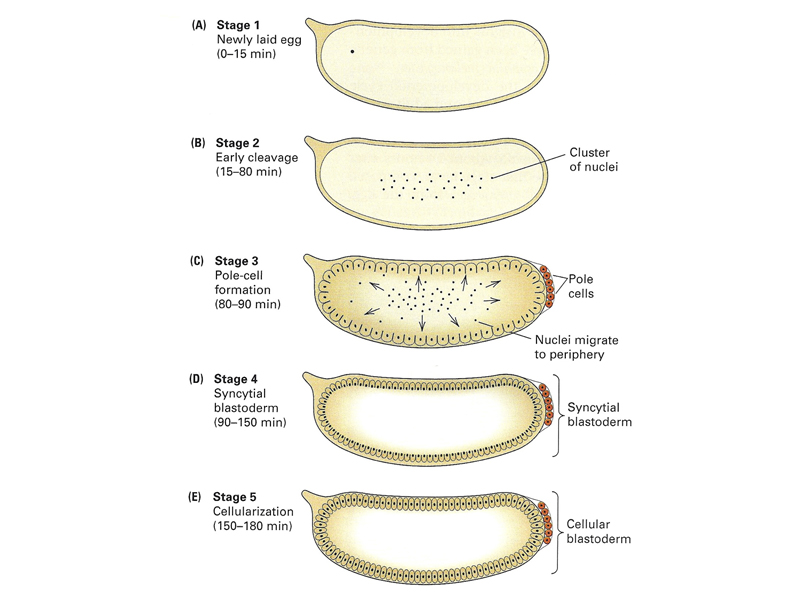
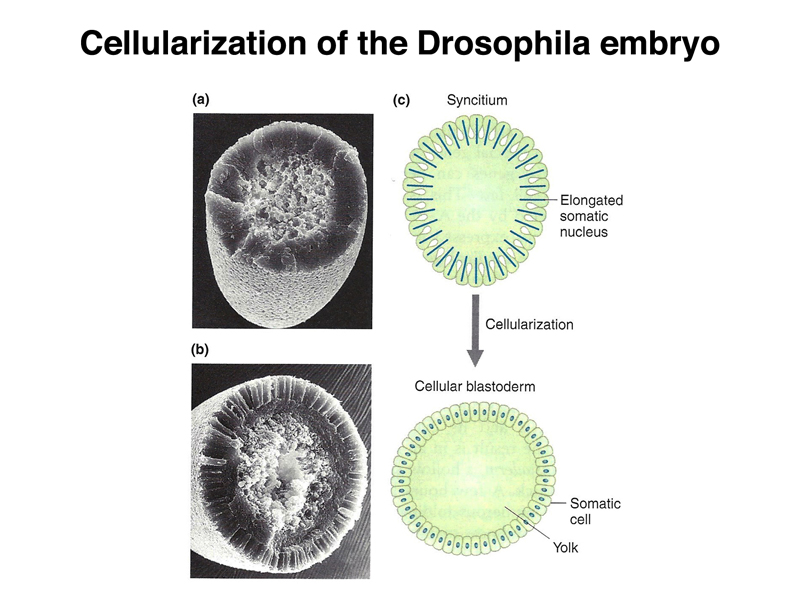
Early Drosophila development is unusual, as shown below. The early nuclear divisions occur rapidly (about every ten minutes) without cell division. Cells then migrate to the periphery. The first nuclei to reach the posterior pole cellularize to become pole cells (the future germ line). Shortly thereafter, all of the other nuclei at the periphery cellulize simultaneously.
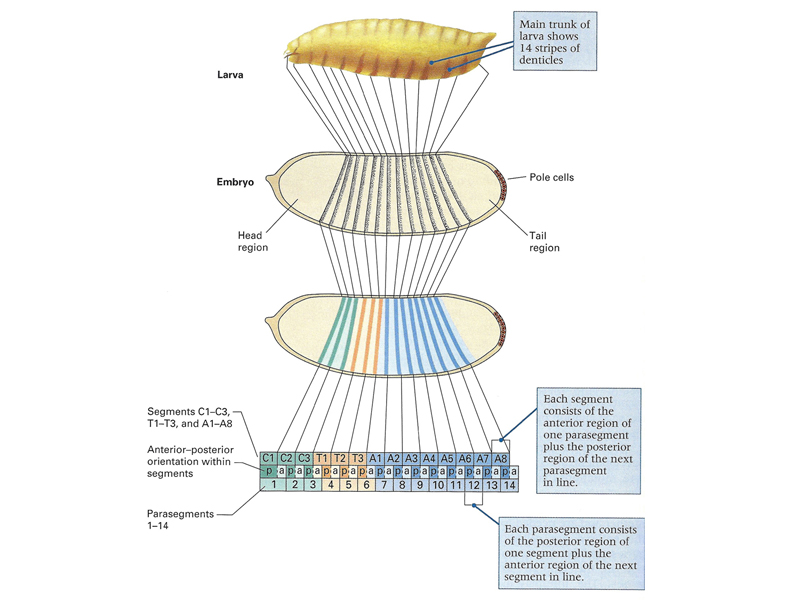
As shown in the figure below, the pattern of the first-instar larva that is the product of embryonic development can be mapped to specific regions of the cellular blastoderm. The larva shows fourteen stripes of denticle belts that reflect the three head, three thoracic, and eight abdominal segments that are evident during embryonic development.
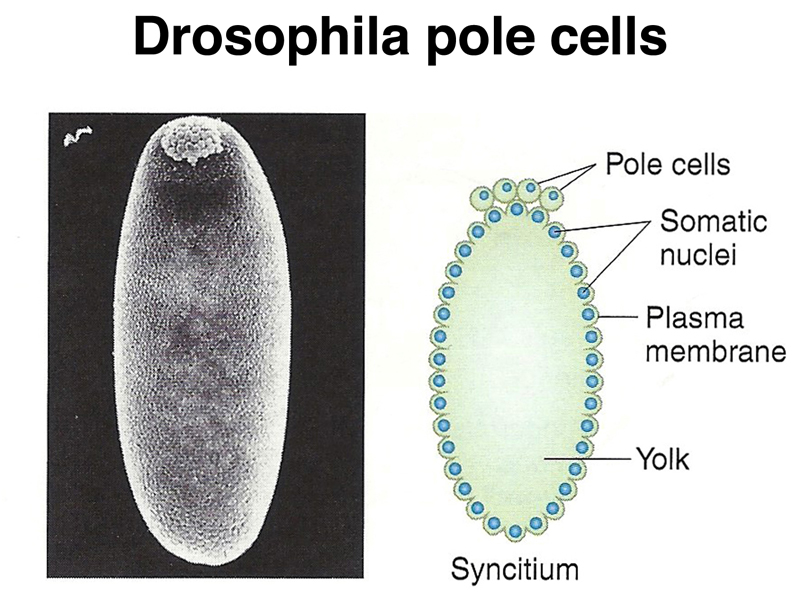
Pole cells, the future germ line, are set aside early in development. The image below shows a scanning electron micrograph of an early embryo with the posterior pole at the top of the image. The pole cells incorporate polar granules, particles of RNA and protein that are deposited in the egg by the mother during oogenesis.
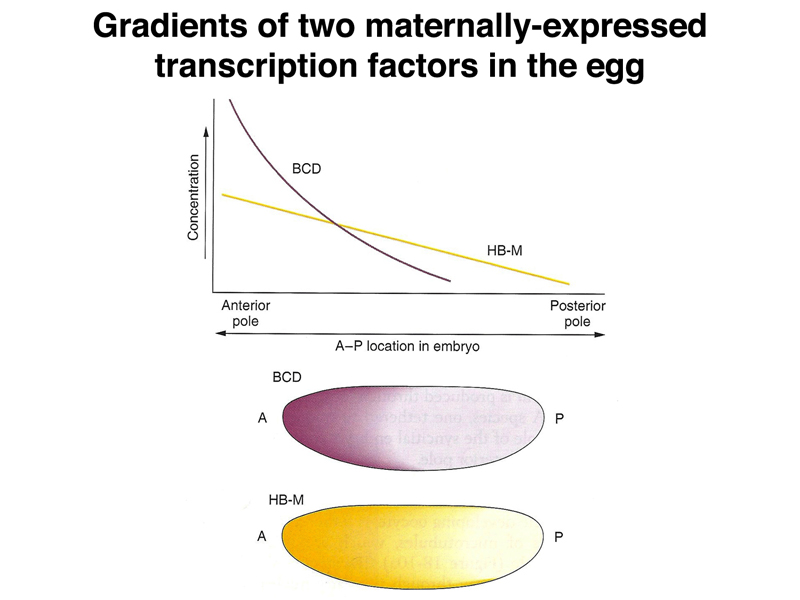
As shown below, there are also localized RNA molecules at the anterior pole of the egg. When these anchored mRNAs are translated, diffusion establishes a gradient of protein along the anterior-posterior axis. The two different protein gradients shown are different, making it possible for cells to read their position by interpreting the concentration of two different proteins. Both proteins are transcription factors.
The figure below shows actual experimental data for two different morphogens, bicoid and nanos. The two panels on the left show in situ hybridization to mRNA. A chemically-labeled nucleic acid probe is made from the cloned bicoid or nanos gene. The probe is denatured and hybridized to a fixed and permeablized embryo. The chemical tag on the probe (for example, biotin) is detected by using a protein that binds the chemical tag (streptavidin in the case of biotin) that is coupled to an enzyme for which there is a chromogenic substrate. The two panels on the right show detection of BCD or NOS protein using antibodies to the protein and secondary antibodies linked to an enzyme for which there is a chromogenic substrate.
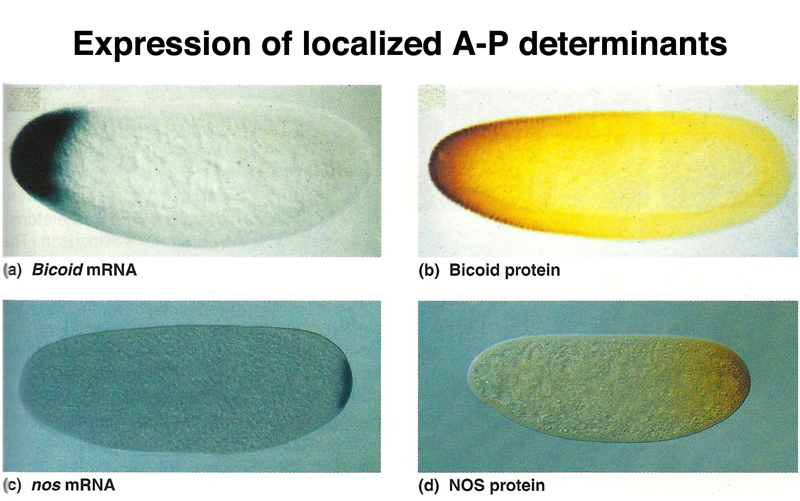
You can see from the figure that the relatively restricted location of the mRNA results in a more broadly distributed gradient of the protein. This tells us that while the mRNA is tethered in some way, the protein product is free to diffuse.
The experiment presented below shows that the signal to localize the bcd mRNA and tether it to the anterior pole is contained within the 3' UTR. A hybrid construct was made that contained the nanos 5'UTR and coding region and the bcd 3' UTR. This hybrid construct, when expressed maternally, localizes to the anterior pole and affects development. The dark-field images that show the larval cuticle show wild-type (left) and mutant (right) phenotypes. In the wild type, we see anterior mouthparts on the left, a series of denticle belts along the ventral surface, and the posterior spiracles on the right. In the mutant, both anterior (left) and posterior (right) show posterior spiracles. The in situ hybridization to RNA shows that while nos mRNA is restricted to the posterior pole in wild type, in the mutant there is nos mRNA at the anterior pole. The distribution of bcd mRNA is not affected.
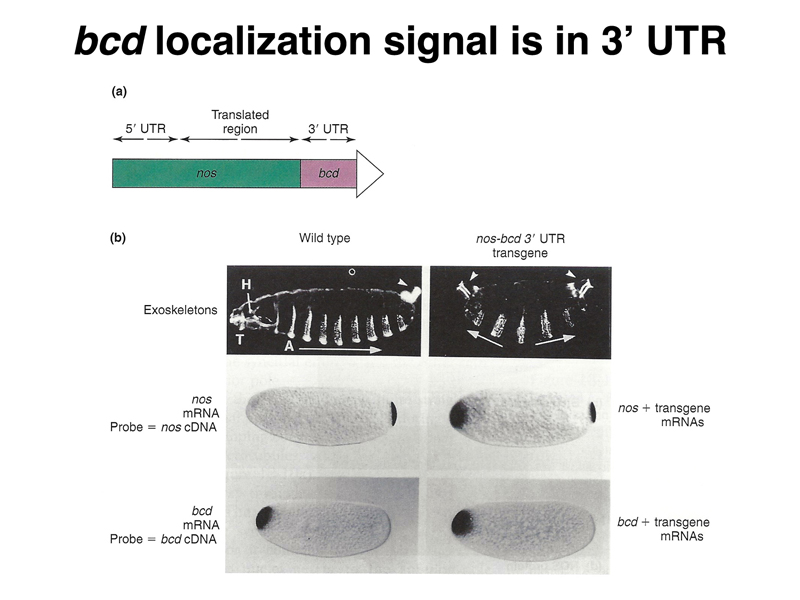
The figure below shows dark-field images of a wild-type larva (top) and a larva from a mother homozygous for bcd (bottom). The embryo from the bcd/bcd mother is missing anterior structures.
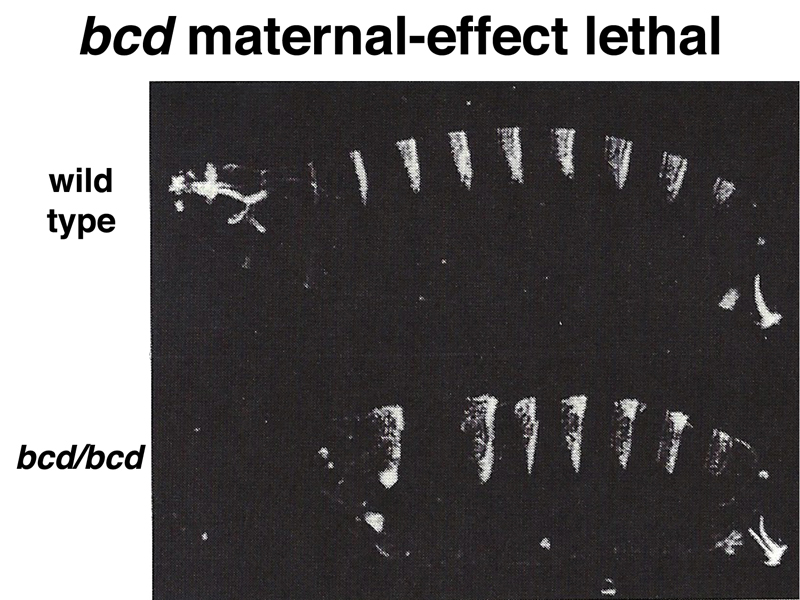
The figure below shows a scanning electron micrograph of a Drosophila embryo that has just formed the cephalic furrow, a cease that marks the boundary between future cephalic and thoracic structures. Pole cells are also visible.
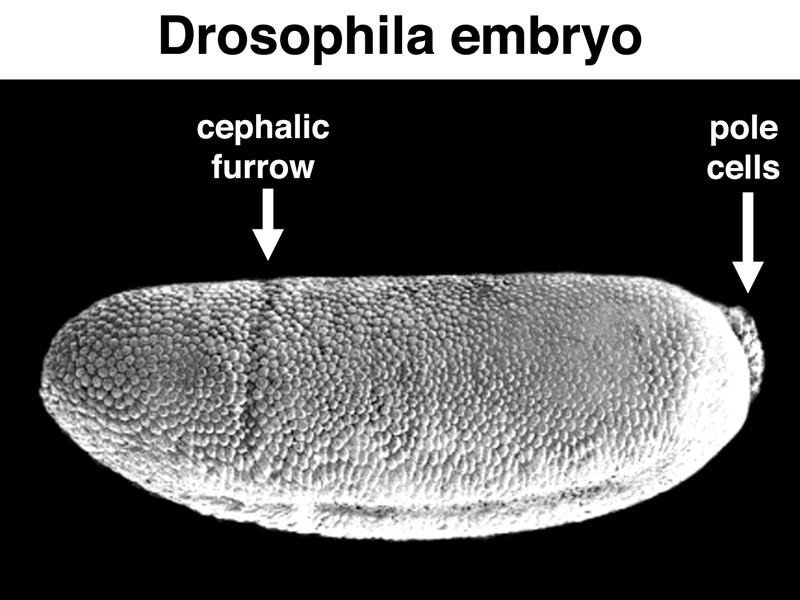
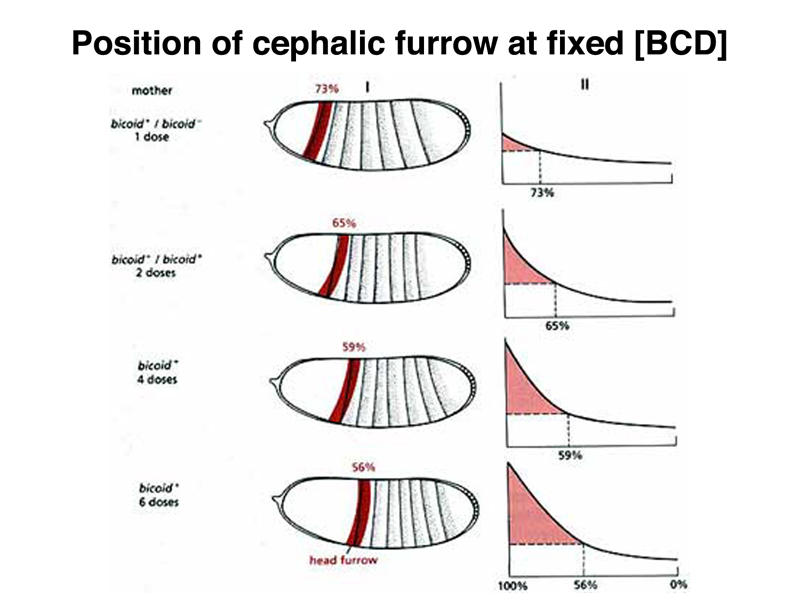
It is possible to vary the dosage of the normal bcd gene in the mother, and hence to cause different amounts of bcd mRNA to be synthesized. This changes the concentration of BCD protein in the early embryo, as shown in the figure below. The position of the cephalic furrow changes together with changes in the distribution of BCD protein.
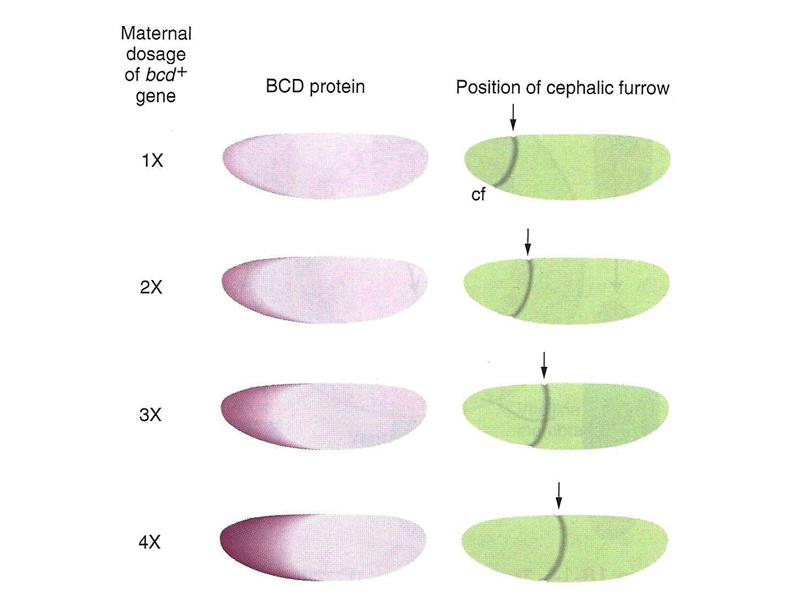
We can interpret these results to mean that there is a particular level of BCD protein that specifies the location of the cephalic furrow, as shown in the figure below.
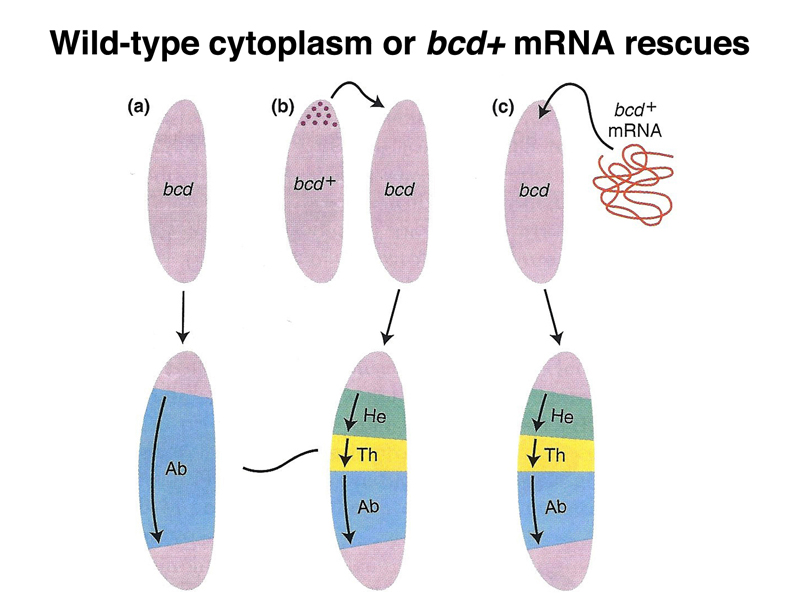
Another experiment shows that anterior cytoplasm of a normal egg is sufficient to rescue an embryo from a bcd/bcd mother, shown below. Transfer of cytoplasm from a normal gee, transfreed by microinjection to an embryo from a bcd/bcd mother will restore the formatin of anterior structures. In addition, bcd mRNA transcribed from a cloned copy of the bcd gene is sufficent to restore anterior structures to an embryo from a bcd/bcd mother.
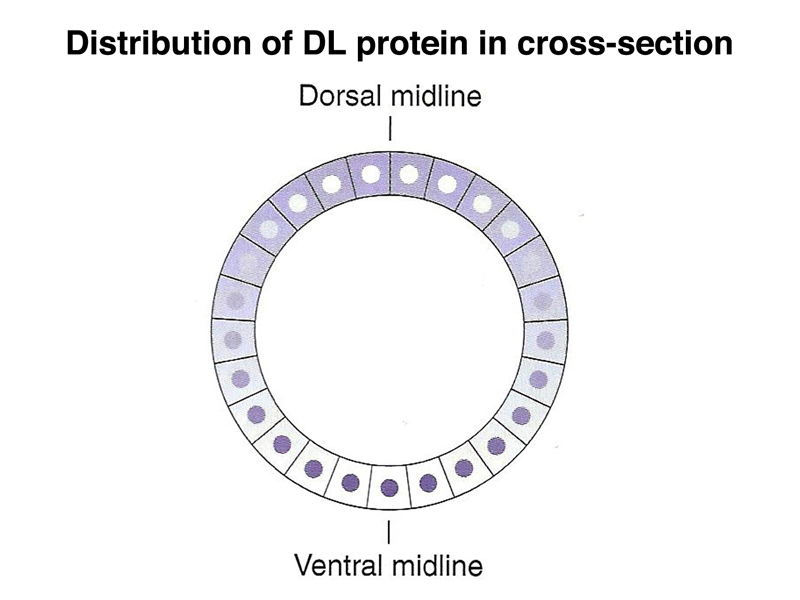
There is also a gradient of a morphogen that specifies the position along the dorsal-ventral axis, as shown below. The DL (dorsal) protein is a transcription factor that is localized to nuclei on the ventral part of the embryo, and to the cytoplasm on the dorsal part of the embryo. There is a concentration gradient of nuclear-localized dorsal protein along the dorsal-ventral axis that helps to specify the position of individual cells.
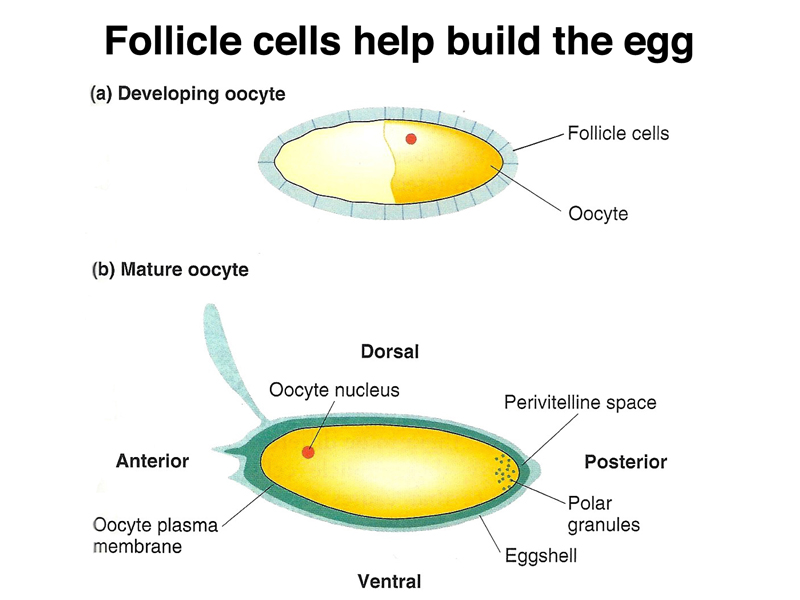
As shown below, during the formation of the egg, follicle cells surround the developing egg, assisting in the formation of the perivitelline space and the eggshell.
At the cellular blastoderm stage, as shown below, there is a concentration gradient of SPZ protein, another morphogen, which is highest at the ventral midline. The SPZ protein binds to the TOLL receptor.
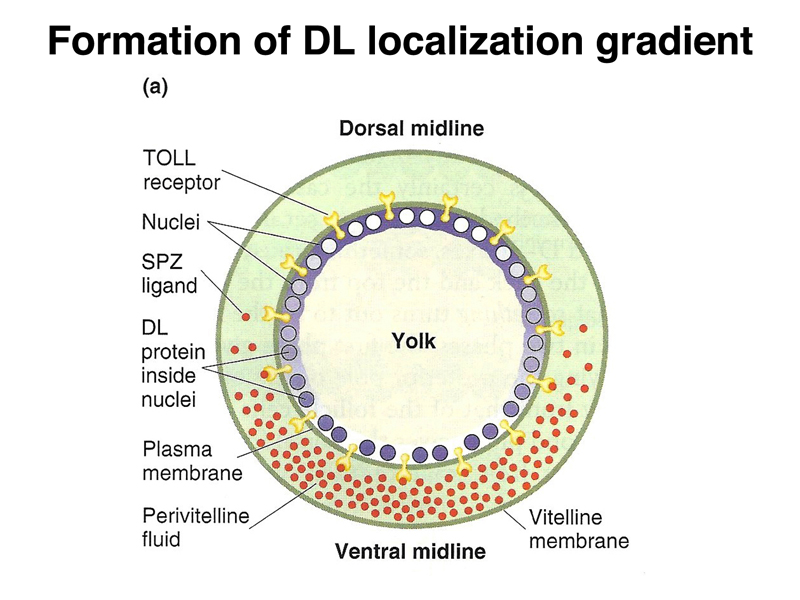
Binding of SPZ to the TOLL receptor leads to the nuclear localization of DL, as shown below. Several other gene products are required. All of these genes were identified by mutations affecting embryonic development.
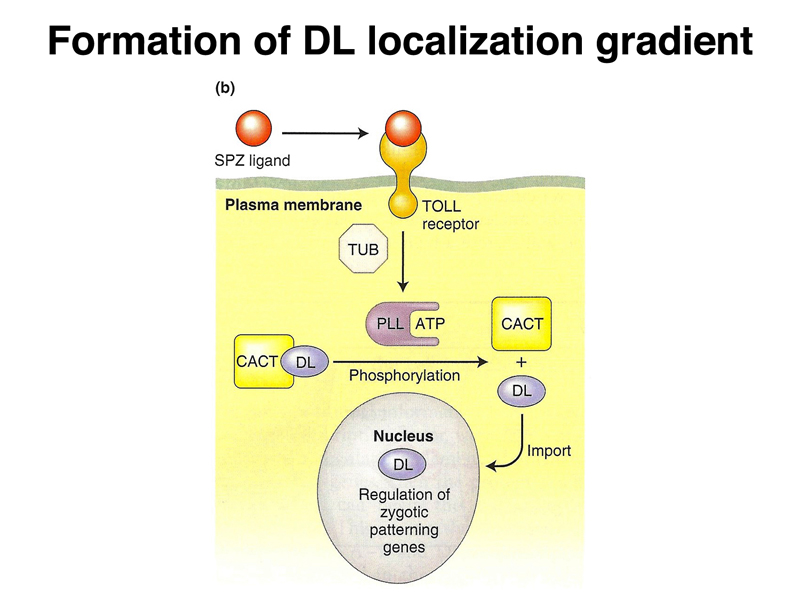
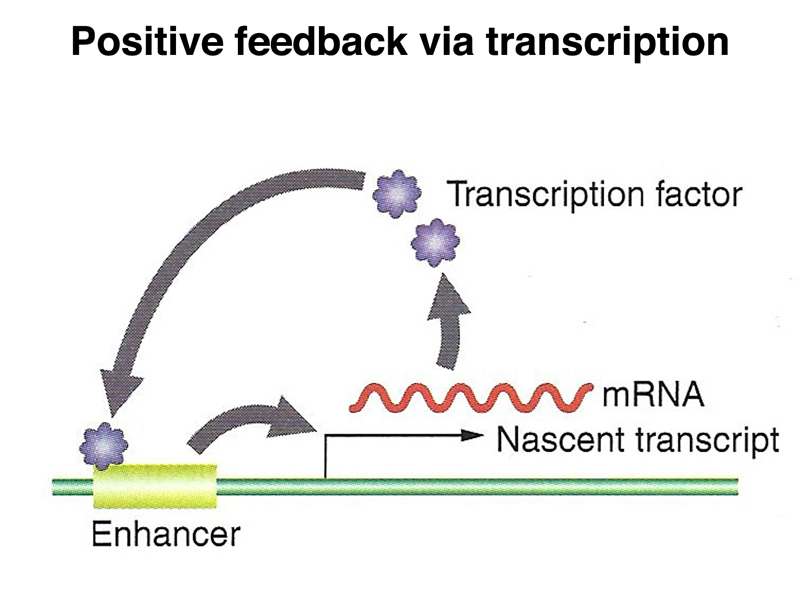
We have seen several examples of a general mechanism in which translation of a tethered mRNA leads to the formation of a concentration gradient of a protein, shown in outline below. The example that we have considered in the greatest detail is bcd.
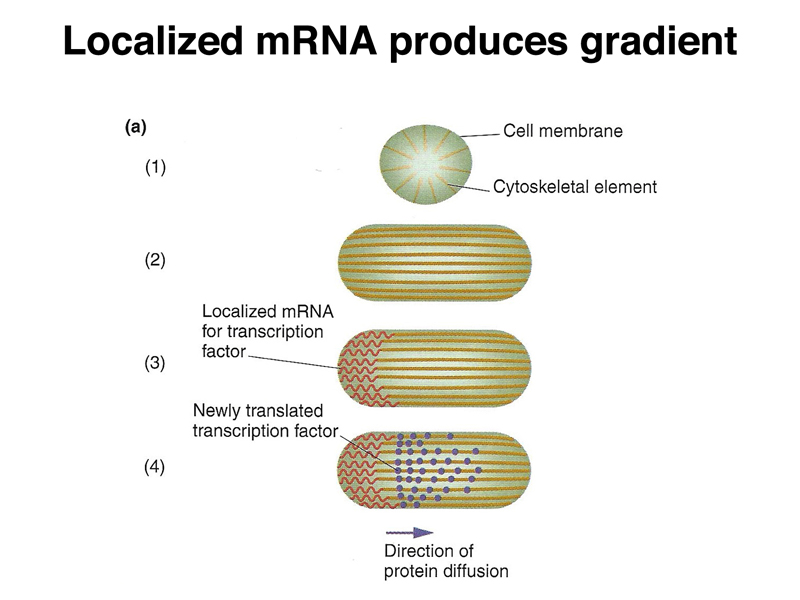
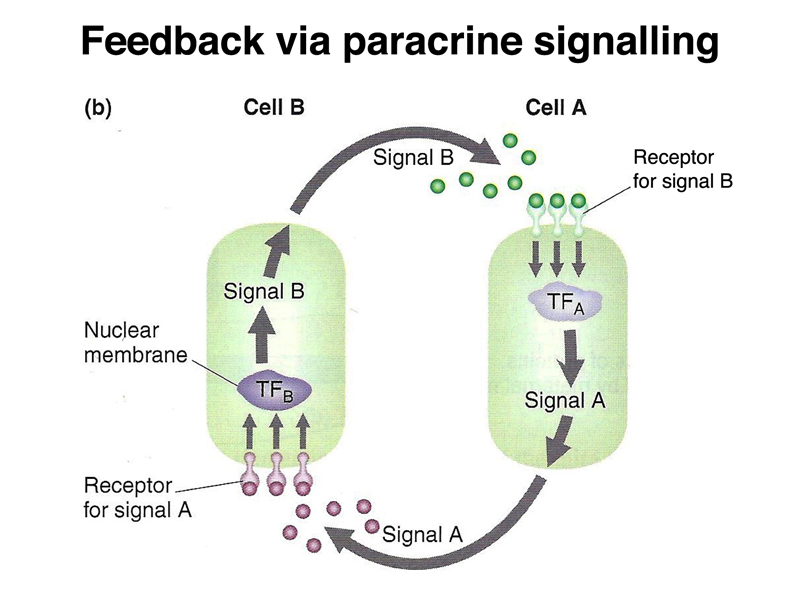
There is another mechanism that is used later in development, shown below. A particular cell can secrete a protein that is a positional-information signal. Neighboring cells, expressing a receptor that recognizes the protein, can interpret their distance from the signalling cell by the concentration of the signal. This is called paracrine signalling.
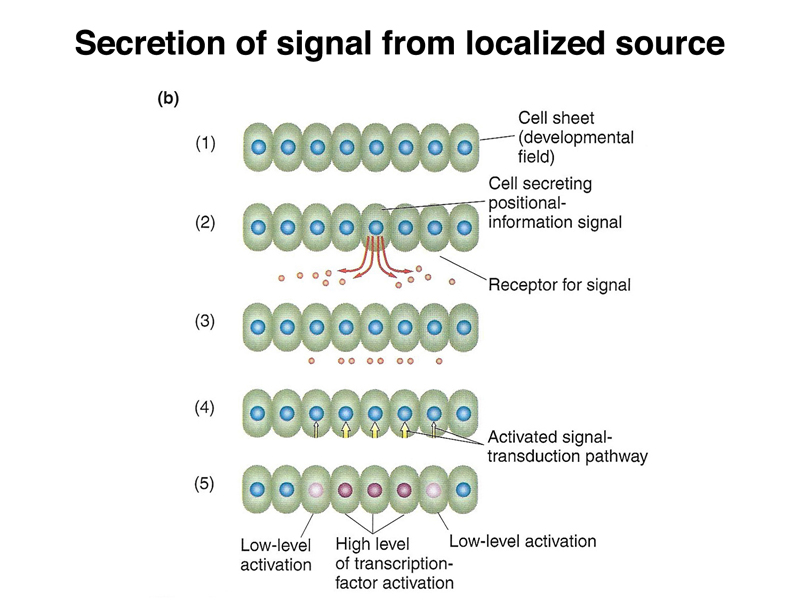
When the Drosophila embryo cellularizes, as shown below, each cell will receive any proteins present in the cytoplasm in that position. In this way, the concentration gradients of morphogen proteins present in the early egg will become differences in the concentration of each morphogen within a cell.
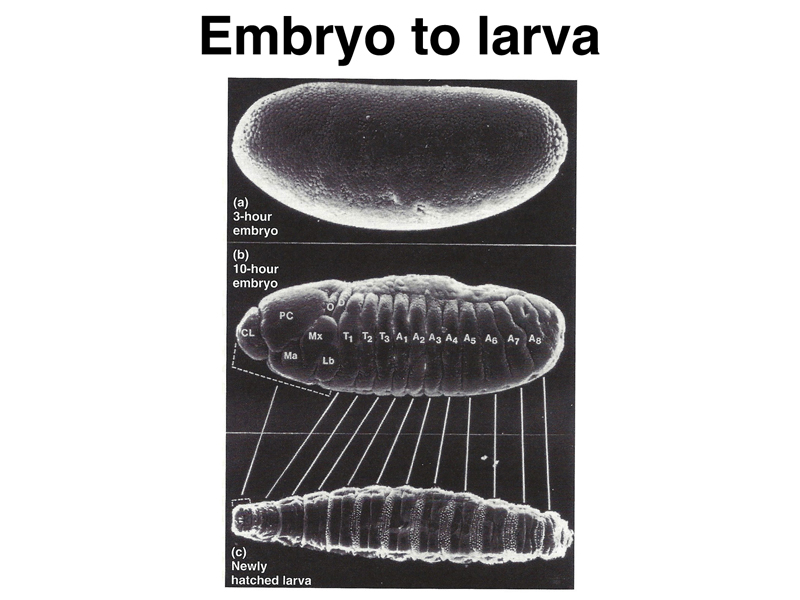
As the embryo develops, we can see a pattern of segmentation that will translate into the segmentation of the first-instar larva. The thoracic (T1 - T3) and abdominal (A1 - A8) segments are evident in the middle image below. The head segments, which occupy a considerable portion of the embryo in the middle image, are less conspicuous in the first-instar larva, due to a process called head involution, kind of like turning a sock partially inside-out.
After the initial events of Drosophila development, leading to the formation of the cellular blastoderm, the first transcription of the zygotic genome occurs. A large screen for recessive mutations that blocked embryonic development identified a number of different kinds of zygotically-active genes. These embryos reach the point of development where they make a larval cuticle, allowing defects in embryonic patterning to be analyzed.
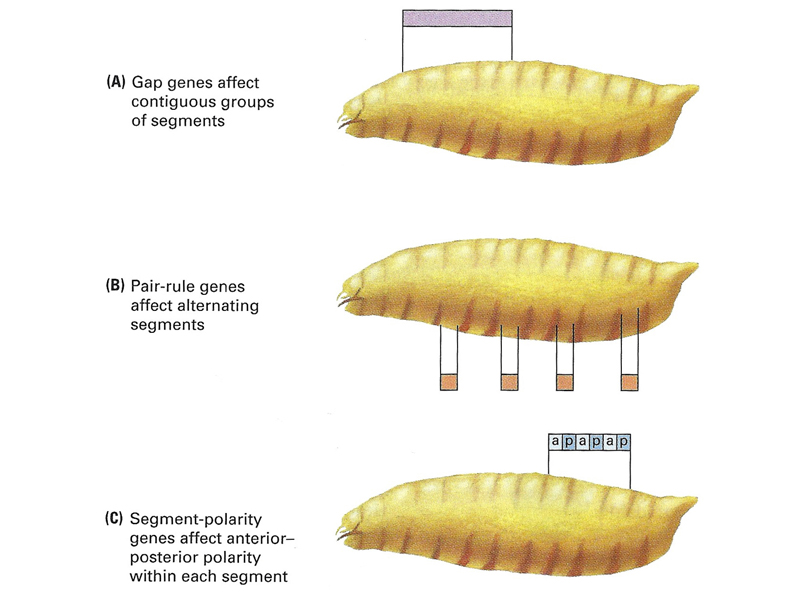
As shown in the figures below, zygotically-acting genes can be placed into three broad categories: Gap genes, pair-rule genes, and segment-polarity genes. The second figure shows the cuticle pattern of defective embryos for a number of zygotic genes, together with an interpretation of which pattern elements are missing, shown as an overlay on a wild-type larva.
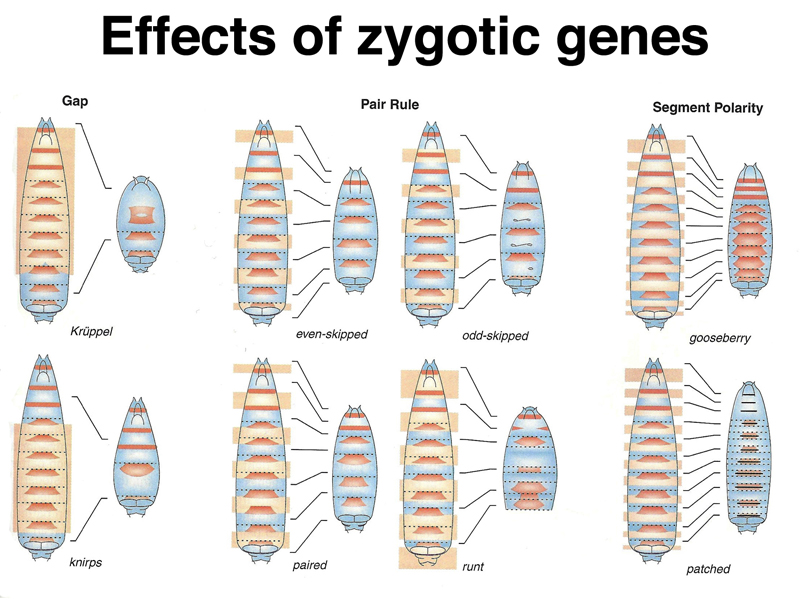
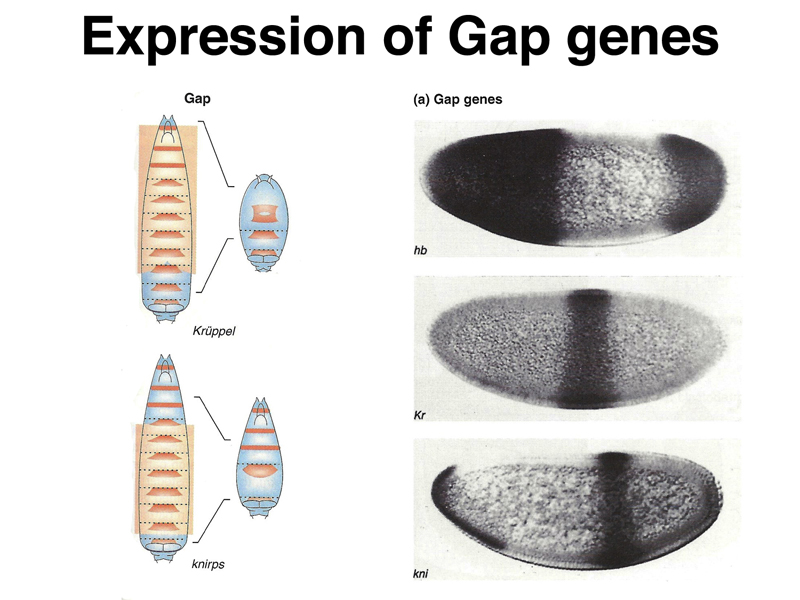
Analysis of the pattern of expression of gap genes, which in mutants are missing several adjacent segments, shows them to be expressed early in broad bands that occupy specific positions along the anterior-posterior axis, as shown below. The photographs on the right show in situ hybridization to mRNA of three different gap genes, hb, Kr, and kni.
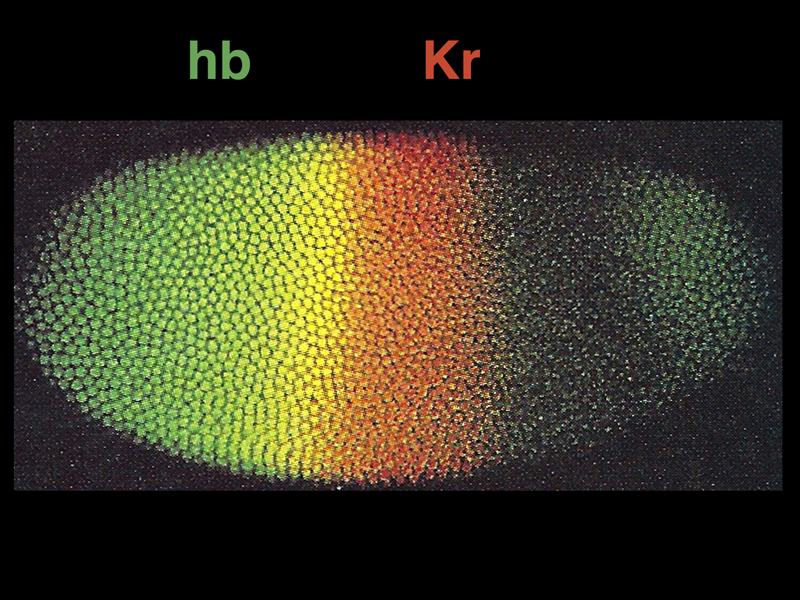
The image below shows antibody detection by immunofluorescence of the protein products of two gap genes: hb (green) and Kr (red). The yellow region shows part of the embryo in which both proteins are present.
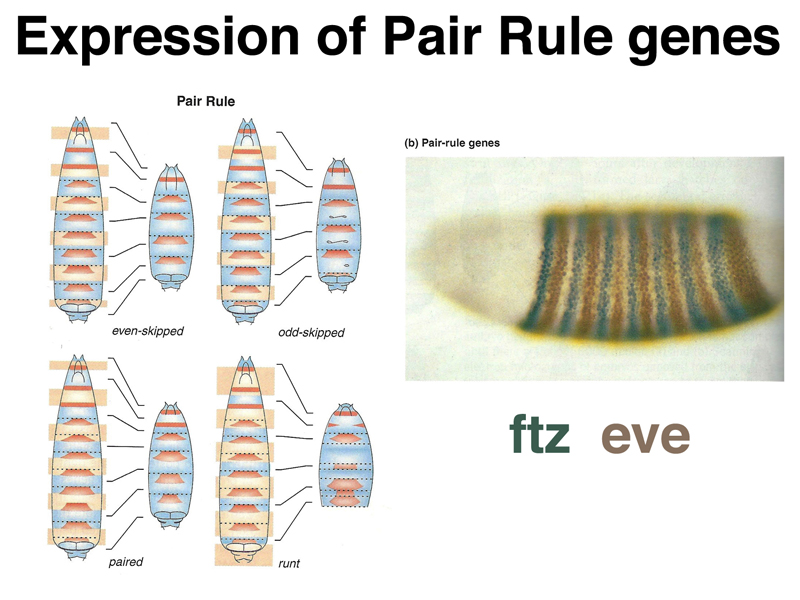
As shown in the image below, pair rule genes create defects in alternating segments in mutants. The photograph on the right shows an embryo stained for the protein products of two different pair rule genes: eve (brown), which is expressed in even-numbered segments, and ftz (blue), which is expressed in odd-numbered segments.
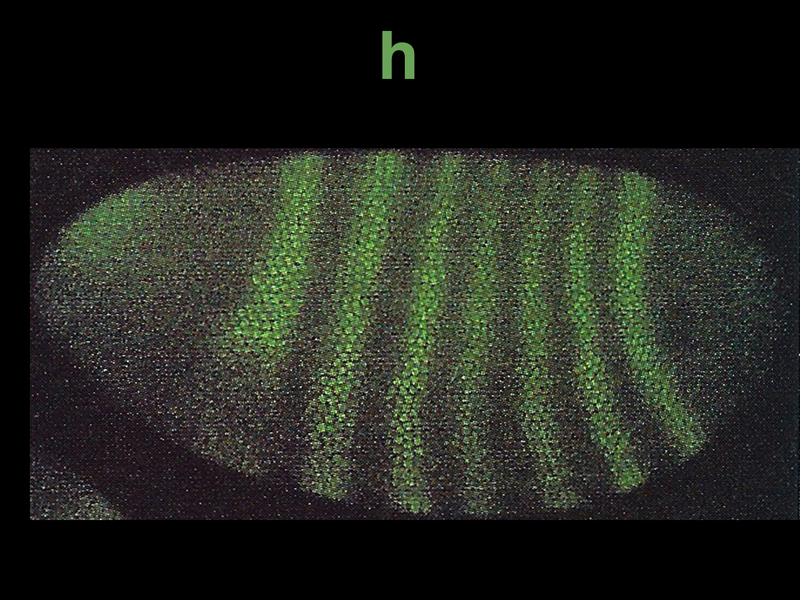
The photograph below shows immunofluorescent staining for h, another pair rule gene. Seven stripes of expression are seen.
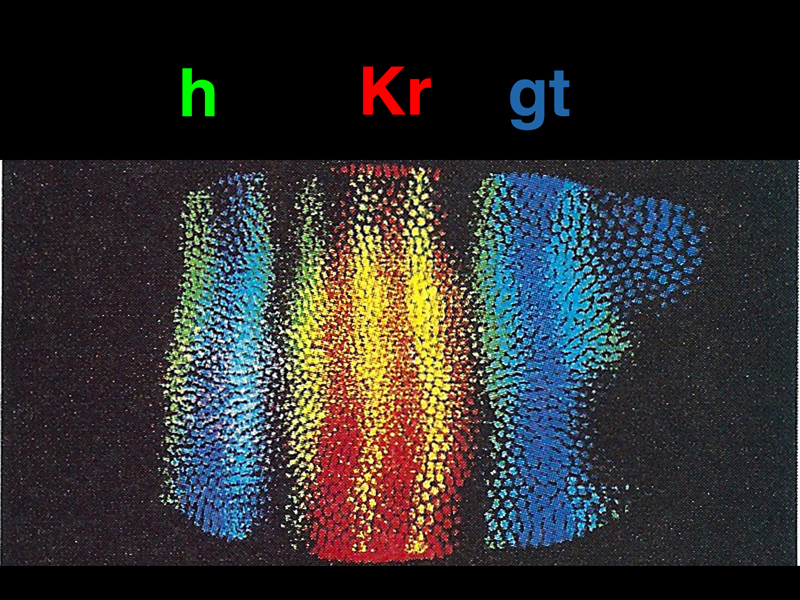
In the image below, we see expression patterns of the protein products of three zygotic genes: Kr and gt (gap genes), and h (a pair rule gene). This shows that the activation of gap and pair rule genes can specify particular locations along the anterior-posterior axis.
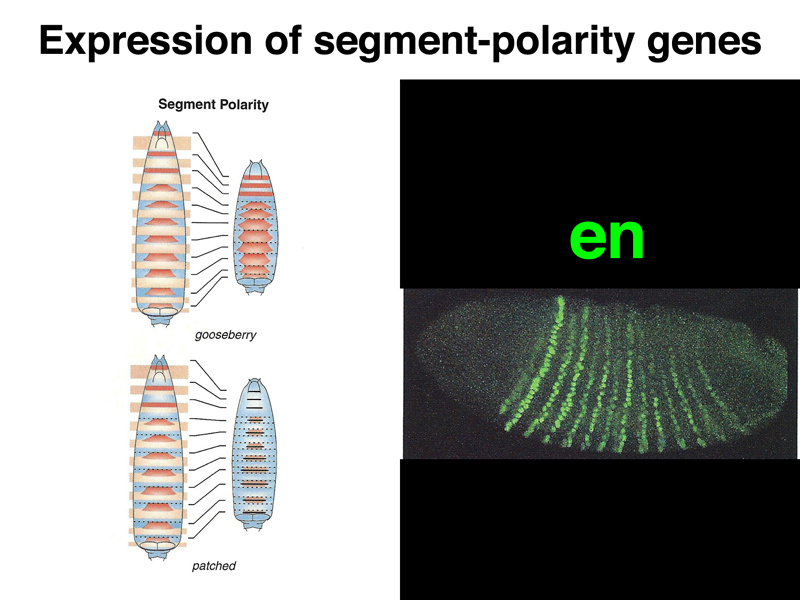
The last zygotic genes to be activated are the segment polarity genes. As shown in the image below, the pair rule gene en is expressed in fourteen stripes that are one or two cell diameters wide each.
Paracrine signalling can also sharpen segment boundaries, as shown below, as cells signal their identities to each other.
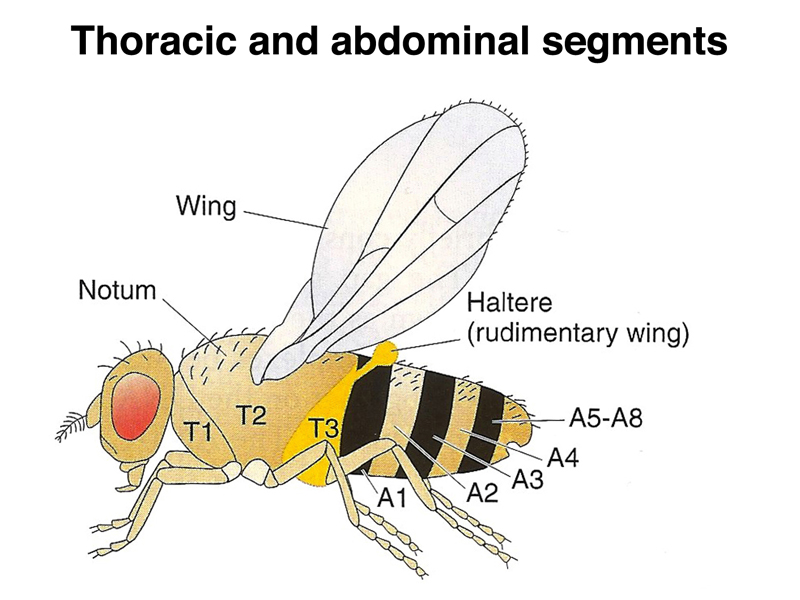
While the gap, pair rule, and segment polarity genes are defining a pattern of segmentation in the embryo, another set of genes is defining the identity of each segment. These are the homeotic genes. A homeotic transformation is the transformation of one body part into another. In order to understand the mutant phenotpye, we need to review the adult anatomy of Drosophila. As in the larvae, there are eight abdominal and three thoracic segments. Unlike primitive marine arthropods, which have both a gill appendage and a leg appendage on each segment, Drosophila has lost all appendages on the abdominal segments. The three thoracic segments each have a leg, while the second thoracic segment has a wing, and the third thoracic segment has a haltere, a kind of vestigial wing. Dragonflies and other winged insects have a two pairs of wings, including a set on the third thoracic segment, but Diptera (two wings) only have wings on the second thoracic segment.
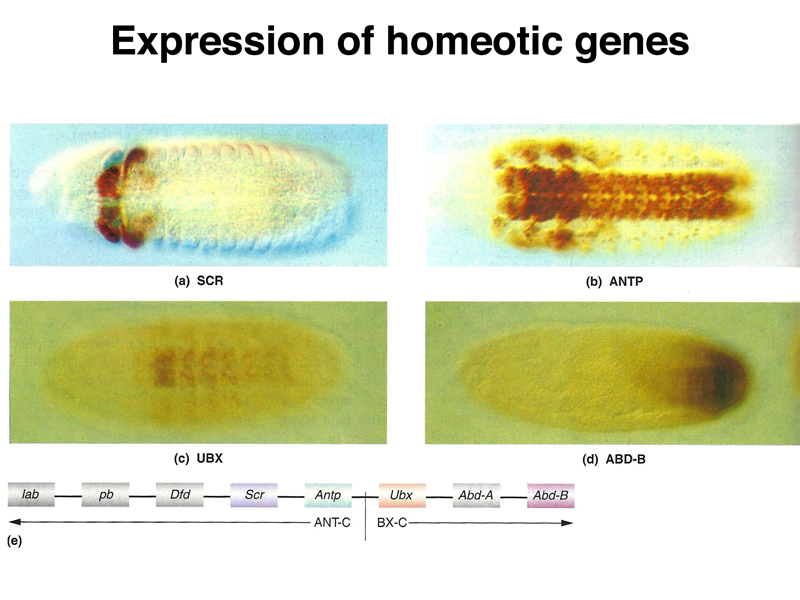
The pattern of expression of several different homeotic genes is shown below. Homeotic genes are expressed across a number of segments. The concentration of the products of particular homeotic genes specifies segmental identity.
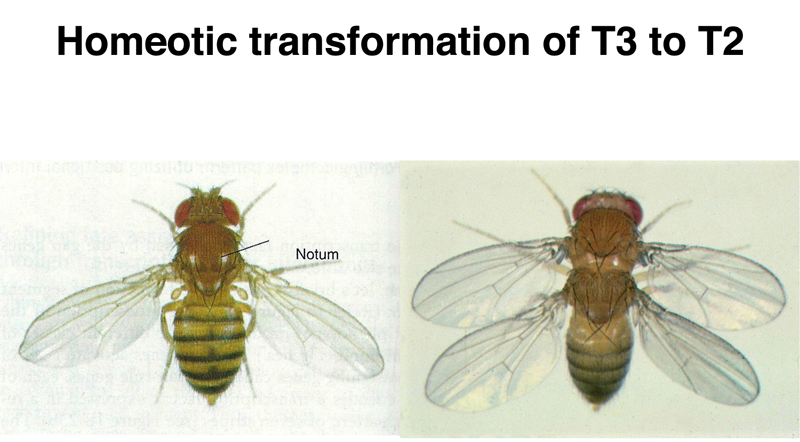
Mutations in homeotic genes produce striking phenotypes. In the figure below, we see a wild-type Drosophila on the left, and a Drosphila transformed by mutations in the bithorax genes on the right. The third thoracic segment is transformed into a second thoracic segment, with a complete set of wings.

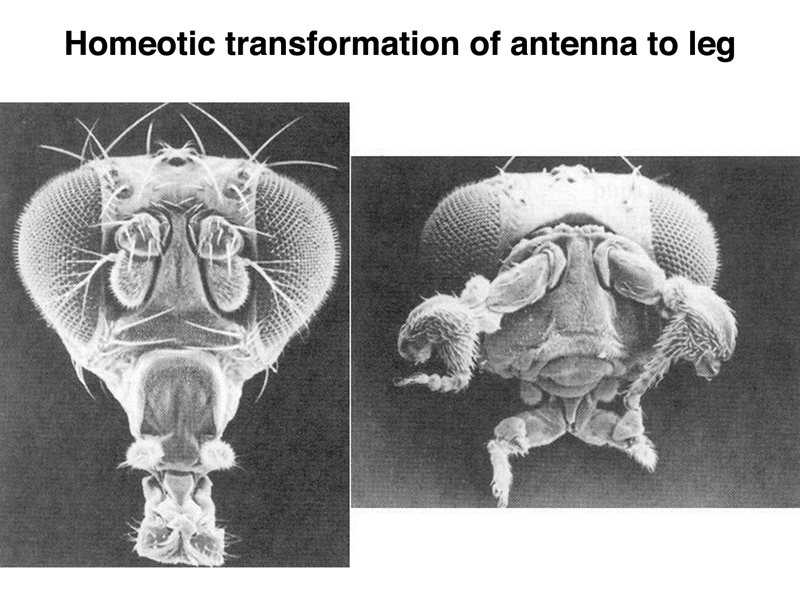
A different homeotic transformation is shown below. A wild-type Drosophila is shown on the left. On the right, a mutation in Antennapedia transforms the antennae to legs.
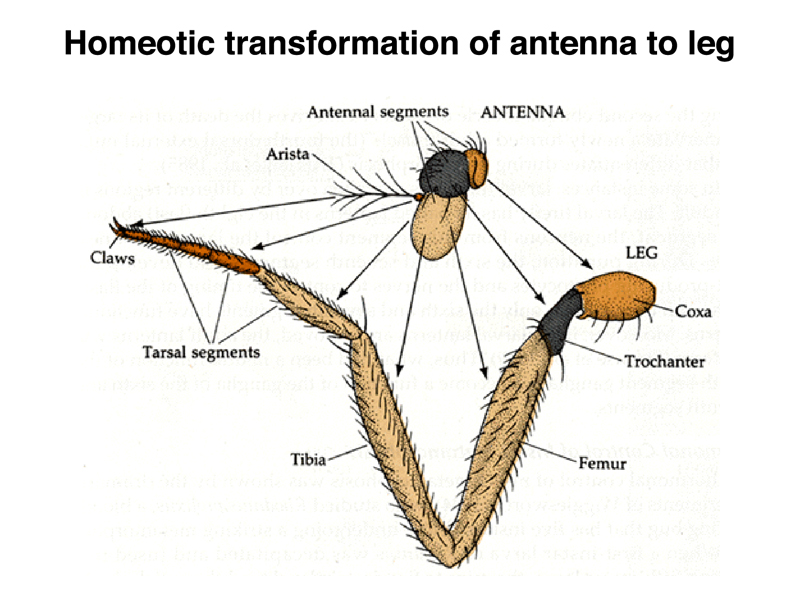
In partial transformations of antenna to leg, we can see that the proximal to distal positional information is preserved, despite the transformation from antenna to leg. Each part of the appendage "knows" where it is along the proximal to distal axis, even if it is transformed to a different appendage.
A student asked what kind of gene can possibly result in such a dramatic transformation. The short answer is that the homeotic genes all encode transcription factors that control the expression of hundreds of other genes.
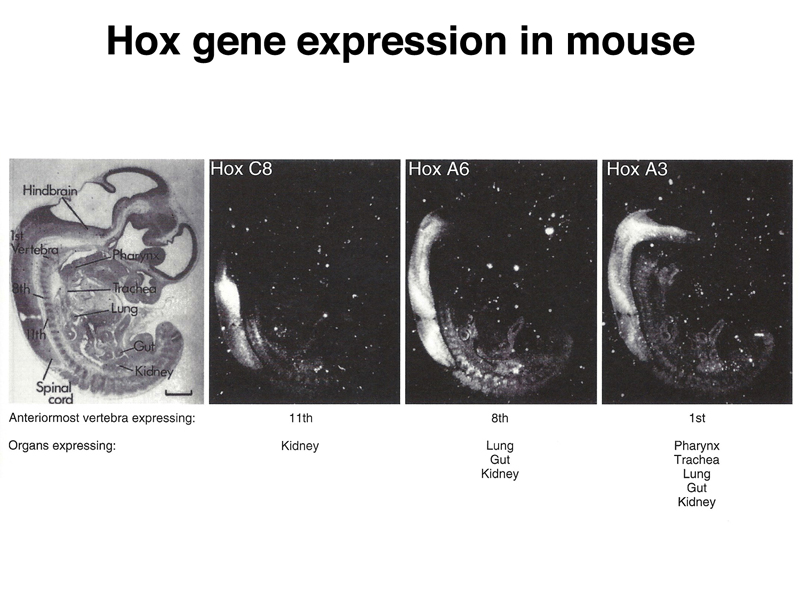
Homeotic genes are conserved in vertebrates, including mice and humans. The figure belwo shows the expression of several Hox (homeobox) genes in mouse embryos. While Drosophila has only a single cluster of Hox genes, mammals have four, reflecting a fourfold duplication of the genome early in vertebrate evolution.
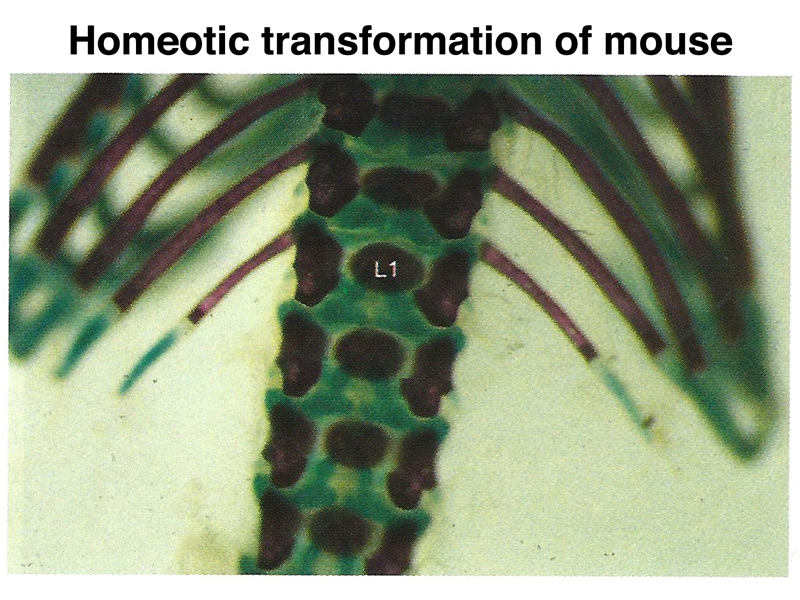
Mutations of specific Hox genes can cause homeotic transformations in the mouse. In the figure shown below, a mouse with a mutation in a Hox gene has a pair of short ribs on a lumbar vertibra, where they are normally absent.
There is another kind of developmental gene resembling a homeotic gene in that it is a transcription factor with hundreds of target genes. The image below shows, on the right, a normal Drosophila and one that is homozygous for a mutation in eyeless, which eliminates eye development. The two photos in the middle show a normal mouse embryo and a mouse embryo homozygous for a loss-of-function allele of the orthologous mouse gene Pax6. The two photos on the left show a normal human eye and the eye of an individual heterozygous for a mutation in Pax6. This causes aniridia, the congenital absence of the iris.
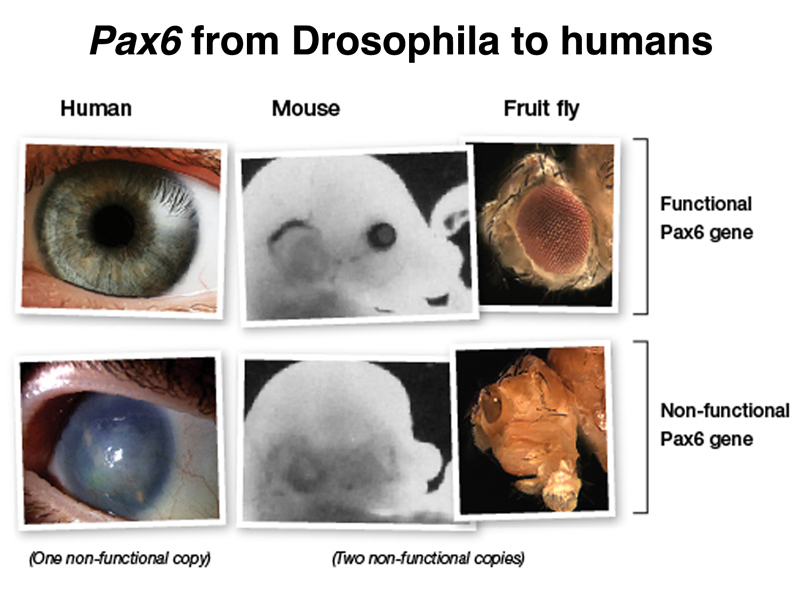
If the human Pax6 gene is expressed in Drosophila homozygous for eyeless, eye development is restored, showing conservation of function of this gene despite the 600 million years of evolution that separate humans from Drosophila.
The figure below summarizes the different types of developmental genes that we have discussed. The coordinate genes are expressed maternally to establish morphogen gradients in the egg. The concentration of various morphogens is interpreted by a set of zygotically-acting gap genes, which have patterns of expression that span several future segments. The positional information is further refined by pair-rule and segment polarity genes that define a pattern of segmentation. Expression of homeotic genes across these segments establishes a set of segmental identities that guides differentiation.