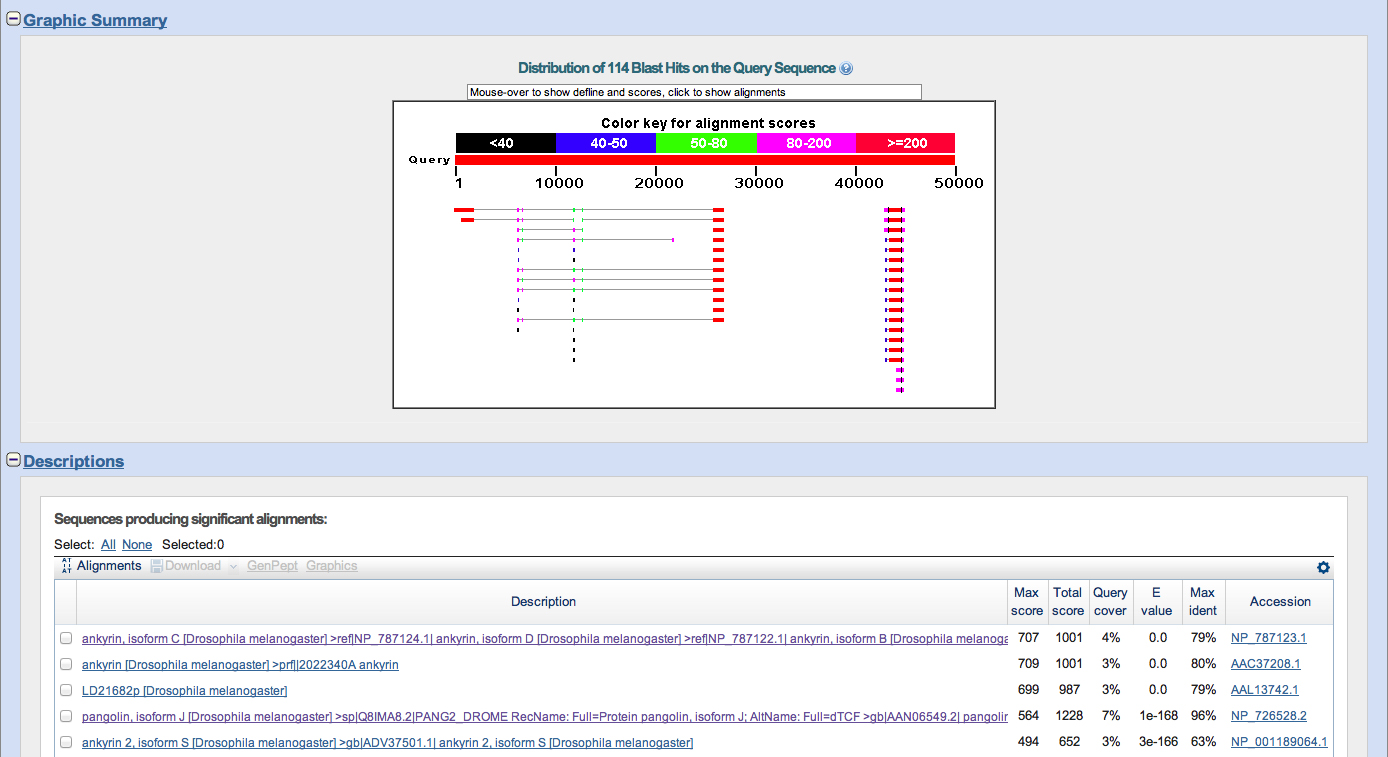
Summary
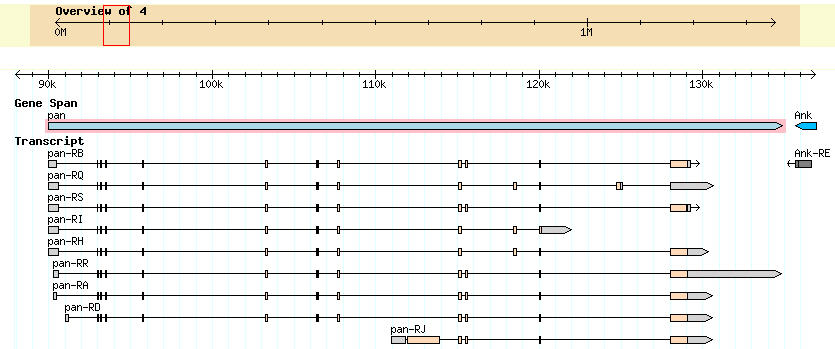
I found 2 genes on contig 5. Pangolin going in the forward direction, and Ankryin in the reverse direction. I do not have the full length of either gene. However, there is overlap between contig 5 and contig 6; contig 6 has the full length of the Ankryin gene. There was some ambiguity as to whether the gene was Ank or Ank2, but is in fact Ank. My GENSCAN analysis confirmed two proteins, both of which checked out. However, there is an exon in the Pangolin gene that was not detected by GENSCAN and is confirmed by the Tophat junction in the Genome browser.

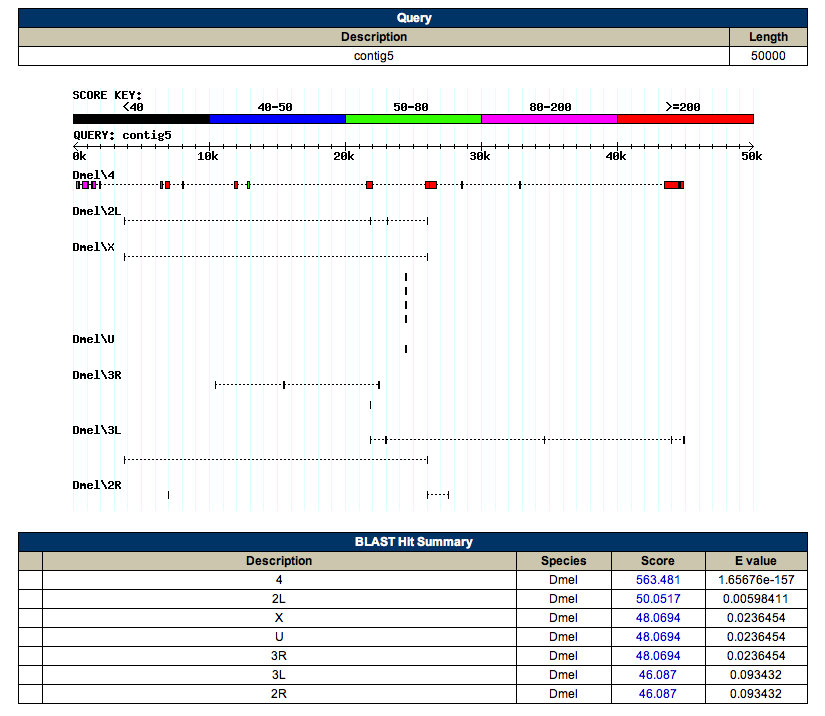
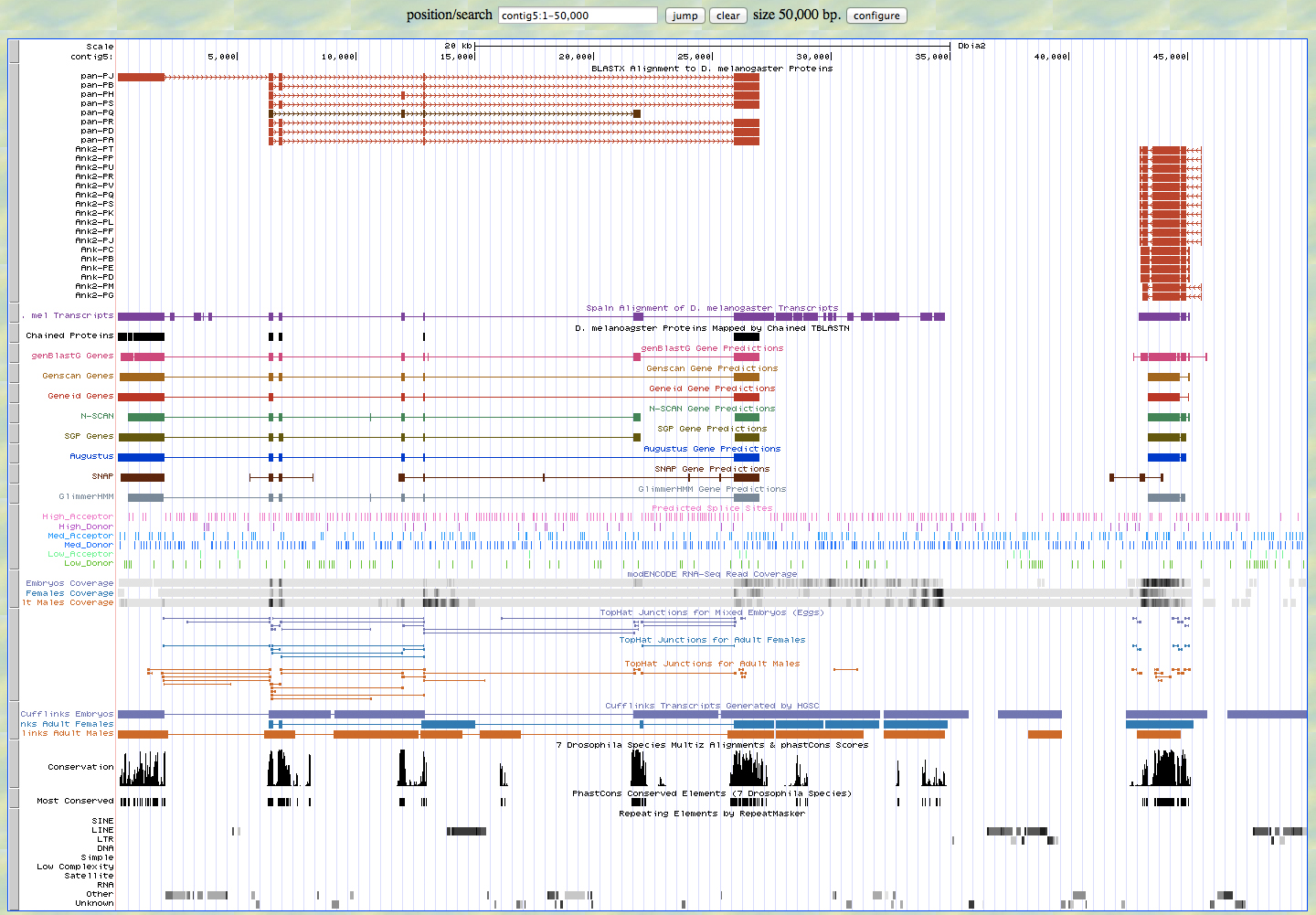
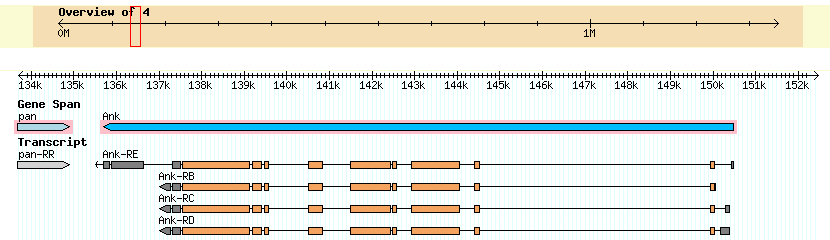
GENSCAN Analysis
>contig5|GENSCAN_predicted_peptide_1|1176_aa VSTNNSTSDDFENIFVVPSRSSAGIASSLSELLLSQHWQNRKTPETDSYQEEVFRNHNEL FSCIYTNSMLNQQKCSEQQLLATQLFYARLLHPHLAEQESFSEKFTDSAANMIHHAELKK PMIGHNELPRTQSSQDNNNNIELTNDLENCSRCGDQPIFKFTKLKKNQKTVQKNMQELEN CNSILNKEQKISQSKESKCTDDYDLQHTCEFIRENKNVLIDVKNRLEKLSVLSAKFRKSL SARQYYVDVNNGSDSALEATVGRQLDGNCKSIESVLEQVKRLYNKWSSAELYYRRSLQRL GLTSEEGVECSPTPTHIIMALAAIALSEEKELSVKETTGIDSKLLEIESVSIKPKTLNEI EDIILQLSSAVNMHQSSVPCTTPDVYSDGVKSDFVETTLIPNCIWHTNKQRFRHKEEGEH CTTAAEIILEYASLSSSSNANTIRVPDTFRSVSNFTDAIENFSVTTQSPNILNNNREGSA VSIISDPSFFPEFSTAPSTPSTSSNSGCSTGIVSGIFGLGHNRRKPRFARRIETQTSDSS KPNVIKESDDQELSNHLLTSSKSTKTANNLCPQILTTTPSIKIAVTTTQERSISNMFEAR LCALSTDTGSLESEIVPMLDAPYDLSIGTKIKPINLDTKNSSNTQSNDAKETSNDKKKPH IKKPLNAFMLYMKEMRAKVVAECTLKESAAINQILGRRWHELSREEQSKYYEKARQERQL HMELYPGWSARDNYGYVSKKKKRKKDRSTTDSGEQAKYYELARRERQLHMQMYPDWSSRT NASRGKKRKRKQDTNDGGNNMKKCRARFGLDQQNQWCKPCRRKKKCIRYMEALNENGPTE DGSGIDEHGSQLSDDDDDDYDDDKLGGSCGSADESNKIGDDDTESLNQSMPSPGCLSGLS SLQSPSTTMSLASPLNMNTNTATNALLIVNADPPTSQQRPTSVSTSGSSSGSTSSISTTP NTSSTVSPVTGTTGPSPSSSHERAMMLGNRFSHLGMGLSPPVVSSSSCNPDTFFQQHPTL CNNAIISLPSIGNSSLHHSSMPNISRNPIGANPRDINNPLSINQLTKRREYQNVELMKDS SSQTIVAHAATSIIHHVAAHGYRANHSLLNSTFSQHFHQQLTNHIETAKMIEPTMLSVST HAVNSTECHKGSDPQANASVKTTSAGASETGVISVS >contig5|GENSCAN_predicted_peptide_2|471_aa XIQHNGIKFIRTSEMELKPDNVPIARPRVQLGPITLEVPHFGTLRENEREIIILRSDDGE SWREHNLYESEGNINDFYKGEHVGDFNQMDELHTDRIVRIVTQNIPHFFAVVSRVRQEVH AIGPDGGTVFSTAVPQVQAIFPPHALTKKIRVGLQAQPVDLVGCSKLLGQGVAVSPIVTV EPRRRKFHKAITLSIPAPKACTKSMVNACYGNGNSNSPTLRLLCSITGGQTRATWEDVTG STPLSFVKDSITFTTTVSARFWLMDCRNVVDAGRMASELYSHLTKVPFLVKFVVFAKRLS KIEAKFSVFCMTDDKEDKTLEQQEYFTEVAKSRDIEVLQDQNIYLEFAGSIVPVLKTGEQ LSTKFQAFCENRLSFIAHIKDPEFPQGRICFMTNPKAGPDELPQNPLCTLNVSVDLKTMT KHLENINESVSLSGNMNYNDLLSSEKRTETAPSEKDIKKLILPEFIRNSAK