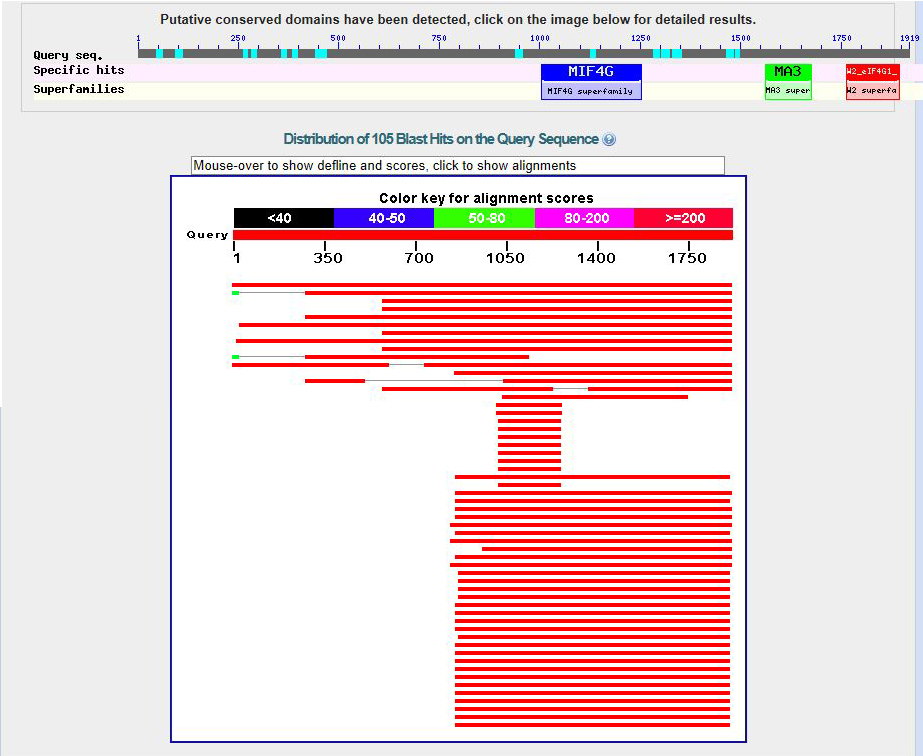
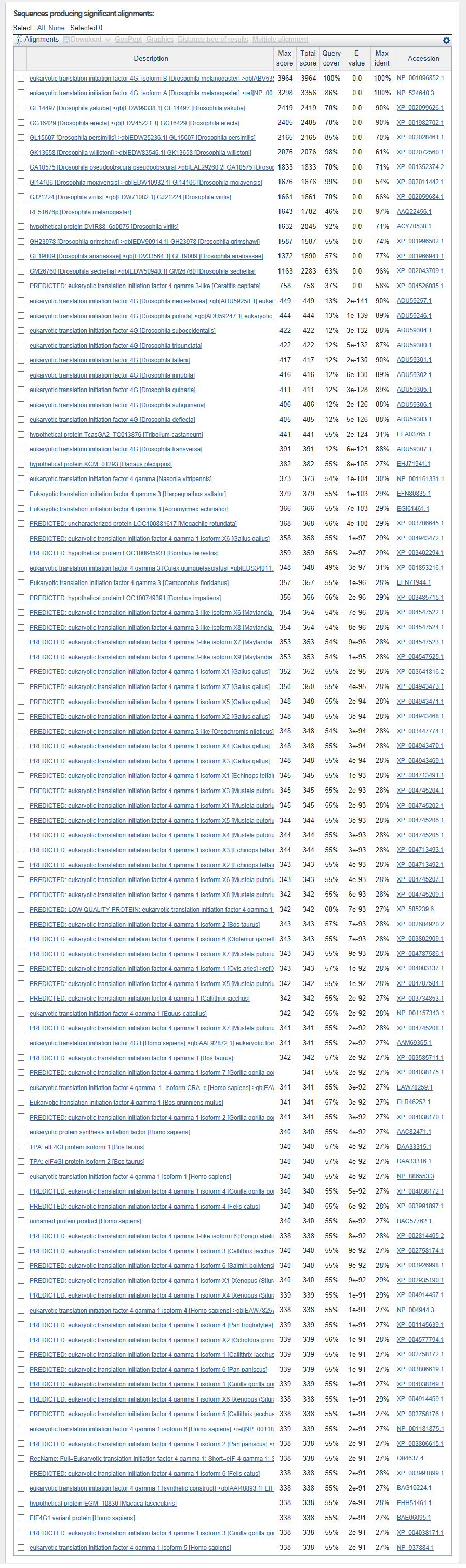
The D. biarmipes eIF4G gene is located on the minus strand of contig50.
The Gene Record Finder gives fifteen CDS segments for D. melanogaster eIF4G:
>eIF4G:15_1629_0
MQQAIPTLPTQSDIDKAMQPHSAQNM
>eIF4G:14_1655_0
ILPANKKTKKYDQQVPTSKPQSLHQPLQPQHSHPTAQPQFQINKAYNVVS
ILKASAQIAQQSPHLTNQQHPPIHHPQQTQQHQQSYTNVVNRSLSASEPV
RAQSSVICNGSSILTVNSRQLNSGDMNSTAIYNISSYRKLTGSLDGNVCF
LNVQDIKQNGNISGSVVSNKSIVGVGSEKSSCTGVSINNQIVLPNAQIGT
SMGIAGTTTAGTSYMHEKNIVGVSVNCVNTSKKYDFNNSSLLSNNSYPAS
TAE
>eIF4G:13_1629_0
YVSTGNNNSGNTRSNPQSGGIFRGPPSTPNAPRGASGGATRHVHVQPMYS
QPLHQNMVLQQYTQYNPRQQTFPASHLQYAPAPMPYYQYQYVPTLQQQPP
HTRSAVTVNTNVNVGNNLQPVQSGPNGPLPVPGASSSQIQLLTSTVQPGA
STVMGVGGPGSTMGQVGVPPMVGVGVMTTSVQPQPVQVQPASRRRHQHRL
QIIDPTTKKNILDDFDK
>eIF4G:12_1629_0
TKSNTDNEFSDQVTSTNTPATVLSEGPIRIPQQESVGLNNLTSTSSQGSE
SRTNAPYIPI
>eIF4G:11_1629_2
TTPISRTDVGPTPIVSAMTDAPSVEILPTPQRGRSK
>eIF4G:10_1629_1
IPIVSPKNVSESIVAPIT
>eIF4G:9_1629_2
ETDDAGSTPISRATEESMPPNQTHPNLLISDSSKHKQAVSNSEISKDAPT
GLKEMHVAELESSVASENHPAGILDVNVNKSSQSLDFSADPEIDSIGAIS
PTSPNAVSPILHEVLDNTELSKKLENSTTERFKDSQSVEKPTHQELSLRN
ATDETEISAMALQDVNSLDNNQIEKTYPSKPKLNVDVSEDISSRESAIKS
TSTKNTGVDVGLQSDSKPETTLNDKQDSTDLKVKVSAKISSIINYNEGQW
SPNNPSGKKQYDREQLLQLREVKASRIQPEVKNVSILPQPNLMPSFIRNN
NNNKRVQSMVGIIGNRSNESAGNYIGKQMSMSGVQSGGGRSSMKGMIHVN
LSLNQDVKLSENENAWRPRVLNKSDGDSDAKSALEKDELVRRVRGILNKL
TPERFDTLVEEIIKLKIDTPDKVDEVIVLVFEKAIDEPNFSVSYARLCQR
LAAEVKVIDERMESETKSNSAHFRNALLDKTEQEFTQNVSQSTAKEKKLQ
PIVDKIKKCTDANEKAELEAFLEEEERKIRRRSGGTVRFIGELFKISMLT
GKIIYSCIDTLLNPHSEDMLECLCKLLTTVGAKFEKTPVNSKDPSRCYSL
EKSITRMQAIASKTDKDGARVSSRVRFMLQDVIDLRKNKWQTSRNEAPKT
MGQIEKEAKNEQLSAQYFGTLSSTTPGGSQGGSGKRDDRGNSRYGESRSS
SAYGGSHSQRGDNGNLRHQQQNNVGGNVSGGAGHSNGNNDENTWHVQTSK
GSRSLAVDSNKLEGL
>eIF4G:8_1629_0
SKLSDQNLETKKMGGLTQFIWISSDTTRLSSAPTPTPSNPFAVLSSLIDK
NSNERDRDRSGPRNKGSYNKGSMERDRYDR
>eIF4G:7_1629_2
MHSRTGSSQGSRENSSSRGGQQGRTLLSSSVQKSTSHSKYTQQAPPTRHT
VK
>eIF4G:6_1629_0
AQSSVGSSNVNTGPLYRGSEQQTSATFSQTTRSVAPVAVFIEASETDLKL
IKSVVSEIVDLSAASKEVTPGAVSCIKRVPEKLRCSFIYYILTDYLHLAN
VGKQYRRYLSIAVSQLIQQNYISADHLRLAYNEFTVYANDLIVDIPELWL
YILQFA
>eIF4G:5_1629_2
PLIVKKILTISDLWNNNLKENSPSNVAKKFLKTYLIYCTQEVGPNFARNM
WIKFNLKWSDFMPESEVADFIKFN
>eIF4G:4_1629_0
RLEYVENESKSPVIDHRETPEKHVKNVIDHIEHLLKEGTTADCIIDYSN
>eIF4G:3_1629_0
GNIMVVDKLFIRGLTETLSNFSIHYKDNSYKLESETFQKFCIPVLQRYID
SNEDHQLECLYTLQLLVHGLEHPR
>eIF4G:2_1629_2
LLSELIGELYDAFVIQKESLCKWRDSKDQSAGK
>eIF4G:1_1629_2
VAVKSLNPFFNSLLNDDAN*
| D. melanogaster eIF4G CDS Segments | BLASTX | Model | Isoform | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| contig coords | alignment | phase | start | end | phase of next exon | A | B | C | |||||
| start | end | frame | E | identity | positive | ||||||||
| eIF4G:1_1629_2 | 16618 | 16559 | -1 | 8e-08 | 95% | 100% | -- | 16620 | 16562 | N/A | X | X | X |
| eIF4G:2_1629_2 | 16780 | 16682 | -1 | 2e-17 | 97% | 100% | 2 | 16782 | 16681 | 2 | X | X | X |
| eIF4G:3_1629_0 | 17139 | 16918 | -2 | 2e-44 | 93% | 98% | 2 | 17139 | 16917 | 2 | X | X | X |
| eIF4G:4_1629_0 | 17346 | 17200 | -2 | 5e-27 | 92% | 97% | 0 | 17346 | 17200 | 0 | X | X | X |
| eIF4G:5_1629_2 | 20394 | 20173 | -2 | 1e-39 | 85% | 94% | 0 | 20396 | 20173 | 0 | X | X | X |
| eIF4G:6_1629_0 | 21072 | 20605 | -2 | 2e-83 | 82% | 90% | 2 | 21078 | 20604 | 2 | X | X | X |
| eIF4G:7_1629_2 | 21397 | 21236 | -1 | 5e-14 | 85% | 90% | 0 | 21399 | 21236 | 0 | X | X | X |
| eIF4G:8_1629_0 | 21699 | 21457 | -2 | 5e-33 | 82% | 89% | 2 | 21699 | 21456 | 2 | X | X | X |
| eIF4G:9_1629_2 | 24624 | 22327 | -2 | 0.0 | 75% | 83% | 0 | 24626 | 22327 | 0 | X | X | X |
| eIF4G:10_1629_1 | 29180 | 29127 | -3 | 2e-04 | 72% | 77% | 2 | 29181 | 29126 | 2 | X | X | X |
| eIF4G:11_1629_2 | 33727 | 33626 | -1 | 7e-16 | 91% | 94% | 1 | 33735 | 33624 | 1 | X | X | X |
| eIF4G:12_1629_0 | 34620 | 34441 | -2 | 7e-24 | 70% | 86% | 2 | 34620 | 34440 | 2 | X | X | X |
| eIF4G:13_1629_0 | 35274 | 34702 | -2 | 2e-44 | 86% | 91% | 0 | 35331 | 34834 | 0 | X | X | X |
| eIF4G:14_1629_0 | 36977 | 36261 | -3 | 1e-58 | 64% | 75% | 0 | 36977 | 36213 | 0 | - | X | - |
| eIF4G:15_1629_0 | 39443 | 39366 | -3 | 4e-10 | 85% | 88% | 0 | 39443 | 39366 | 0 | X | X | X |

CDS13 is in frame -2 and has three possible donor sites. At base 34675, there is BLASTX and RNA-seq support, but there is an in-frame stop codon before the splice site. At base 34768, there is very little evidence of a splice site; only one gene predictor supports this site. At base 34834, there is support from gene predictors and splice predictors. All splice sites are in phase 0. A splice site at 34834 would require a deletion of 90 bases from D. melanogaster, so I compared the gene to other species to determine how well conserved the region is.